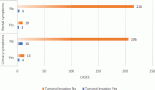
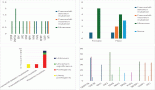
A wide range of microbes inhabit the oral cavity, and bacterial and fungal communities most often exist as structured communities or biofilms. The use of tobacco alters the structure of the oral microbiome, including that of potentially malignant lesions, and the altered oral microbiome influences key microenvironmental changes such as chronic inflammation, secretion of carcinogenic toxins, cellular and tissue remodelling and suppression of apoptosis. Given this, it is clear that the bacterial and fungal biofilms in potentially malignant states are likely not passive entities, but could play a critical role in shaping potential malignant and carcinogenic conditions. This holds potential towards leveraging the oral microbiome for the management of tobacco-associated potentially malignant lesions and oral cancer. Here, we explore this line of investigation by reviewing the effects of tobacco in shaping the oral microbiome, and analyse the available evidence in the light of the microbiome of oral potentially malignant and cancerous lesions, and the role of dysbiosis in carcinogenesis. Finally, we discuss possible interventions and approaches using which the oral microbiome could be leveraged towards precision-based oral cancer therapeutics.